Spatio-temporal dbscan with network distance
Jeremy Gelb
2025-10-04
Source:vignettes/web_vignettes/SpaceTimeDBscan.Rmd
SpaceTimeDBscan.RmdIntroduction
This vignette is just a simple example about what can be done with
spNetwork and little bit of imagination. We will performe
here a spatio-temporal dbscan clustering with network distances on the
bike_accident dataset. Here is the procedure:
- calculating a network distance matrix between each pair of points;
- calculating a temporal distance matrix between each pair of points;
- Calculating a binary matrix as the intersection of the two previous matrices according to a temporal and a network maximum distance thresholds;
- using the dbscan algorithm on the binary matrix;
- analysing the results.
Calculating the two distance matrix
This is the first step, we will start with the network matrix.
# first load data and packages
library(sf)
library(spNetwork)
library(spdep)
library(dbscan)
library(tmap)
data(mtl_network)
data(bike_accidents)
distance_mat_listw <- network_listw(bike_accidents, mtl_network,
maxdistance = 5000,
dist_func = "identity",
matrice_type = "I",
grid_shape = c(1,1))
# spdep changed its name rom nb_list to neighbours
distance_mat_listw$neighbours <- distance_mat_listw$nb_list
distance_mat_net <- listw2mat(distance_mat_listw)Great, now we will calculate a temporal distance matrix in days between the accidents.
bike_accidents$dt <- as.POSIXct(bike_accidents$Date, format = "%Y/%m/%d")
start_time <- min(bike_accidents$dt)
bike_accidents$time <- difftime(bike_accidents$dt,start_time, units = "day")
temporal_mat <- as.matrix(dist(bike_accidents$time))Calculating the intersection matrix
We select here the two following thresholds: 500m and 25 days. We calculate a binary matrix indicating if two points are close enough in time and space to belong to the same cluster.
binary_mat <- as.integer(temporal_mat <= 25 & distance_mat_net<= 400)
dim(binary_mat) <- dim(temporal_mat)Applying the dbscan algorithm
The last step is to just apply the dbscan algorithm!
result <- dbscan(binary_mat, eps = 1, minPts = 5)
result
#> DBSCAN clustering for 347 objects.
#> Parameters: eps = 1, minPts = 5
#> Using euclidean distances and borderpoints = TRUE
#> The clustering contains 4 cluster(s) and 325 noise points.
#>
#> 0 1 2 3 4
#> 325 7 5 5 5
#>
#> Available fields: cluster, eps, minPts, dist, borderPointsAnalysing the results
We will first map the clusters (everybody loves map).
tmap_mode("view")
bike_accidents$cluster <- as.character(result$cluster)
out_cluster <- subset(bike_accidents,bike_accidents$cluster == "0")
in_cluster <- subset(bike_accidents,bike_accidents$cluster != "0")
tm_shape(out_cluster) +
tm_dots("black", alpha = 0.3, size = 0.01) +
tm_shape(in_cluster) +
tm_dots("cluster")And finally, we will plot the time period of each cluster.
ggplot(in_cluster) +
geom_point(aes(x = dt, y = cluster, color = cluster)) +
scale_x_datetime(date_labels = "%Y/%m")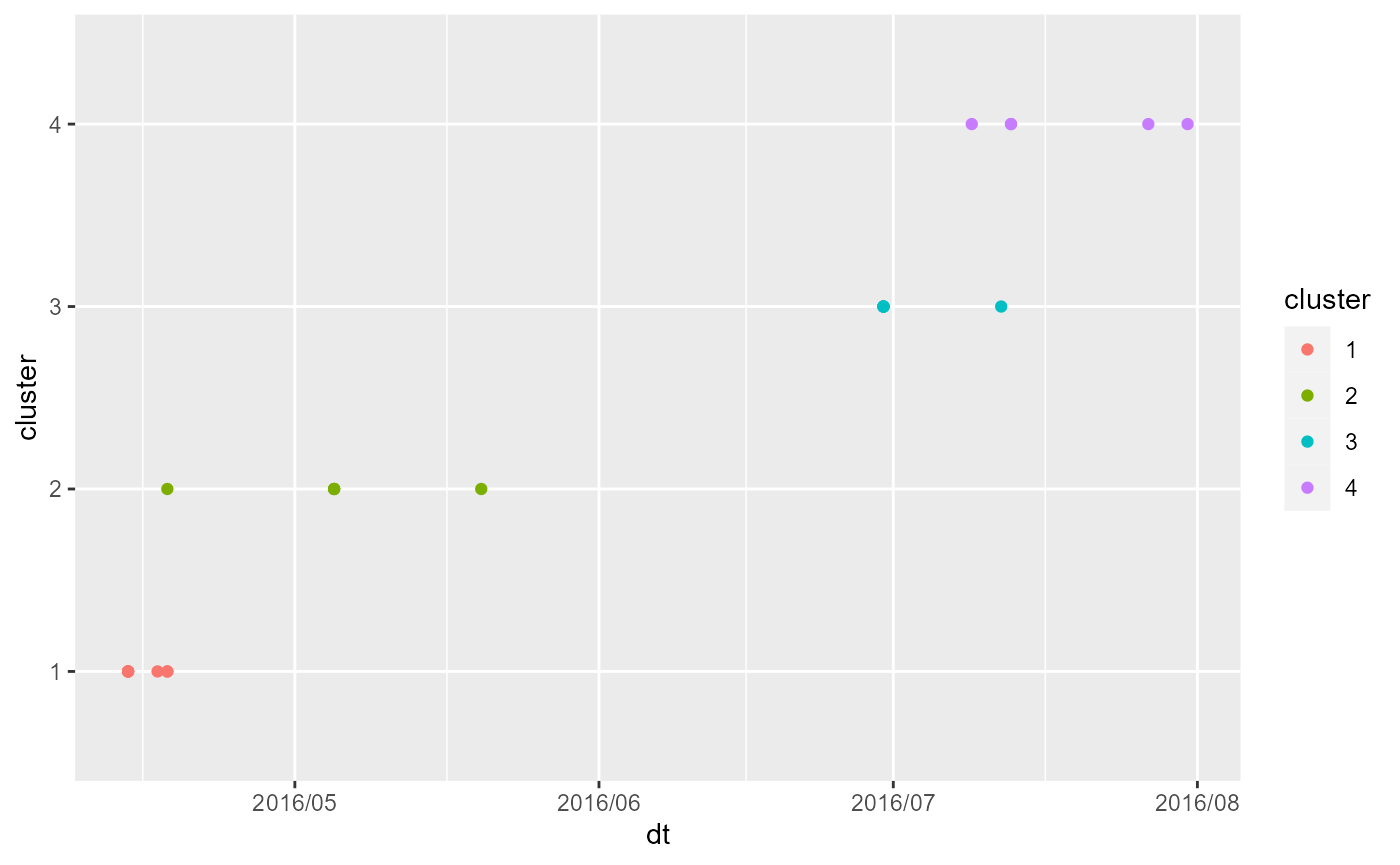
That’s all folks! I hope this short example was interesting!